1. Load the package
Open grayleafspotr/grayleafspotr.Rproj in RStudio for
the source workflow, or install the package and load it with
library(grayleafspotr).
2. Confirm the example data
example_dir <- system.file("extdata", "example", package = "grayleafspotr")
list.files(example_dir)## [1] "analysis.csv" "analysis.json" "manifest.json"3. Load saved example results
example_run <- example_grayleafspot_results()
example_run$run## $id
## [1] "example-run"
##
## $engine
## [1] "python-local"
##
## $engineModel
## [1] "packaged-python-pipeline"
##
## $createdAt
## [1] "2026-04-14T12:00:00Z"
##
## $outputDir
## [1] "inst/extdata/example"
##
## $analysisJson
## [1] "inst/extdata/example/analysis.json"
##
## $analysisCsv
## [1] "inst/extdata/example/analysis.csv"
example_run$results## # A tibble: 2 × 34
## id filename day area_mm2 radius_mm diameter_mm perimeter_mm circularity
## <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 demo-1 demo_day… 1 12 1.95 3.9 8 0.8
## 2 demo-2 demo_day… 2 18 2.39 4.78 9.1 0.77
## # ℹ 26 more variables: eccentricity <dbl>, edge_roughness <dbl>,
## # contrast <dbl>, correlation <dbl>, energy <dbl>, homogeneity <dbl>,
## # entropy <dbl>, center_edge_delta <dbl>, density_index <dbl>, core <dbl>,
## # middle <dbl>, outer <dbl>, crack_count <dbl>, crack_length_mm <dbl>,
## # crack_coverage_pct <dbl>, proportional_crack_coverage_pct <dbl>,
## # radial_velocity_mm_per_day <dbl>, area_growth_rate_mm2_per_day <dbl>,
## # relative_growth_rate_per_day <dbl>, radial_acceleration <dbl>, …4. Generate template plots
plot_colony_expansion(example_run)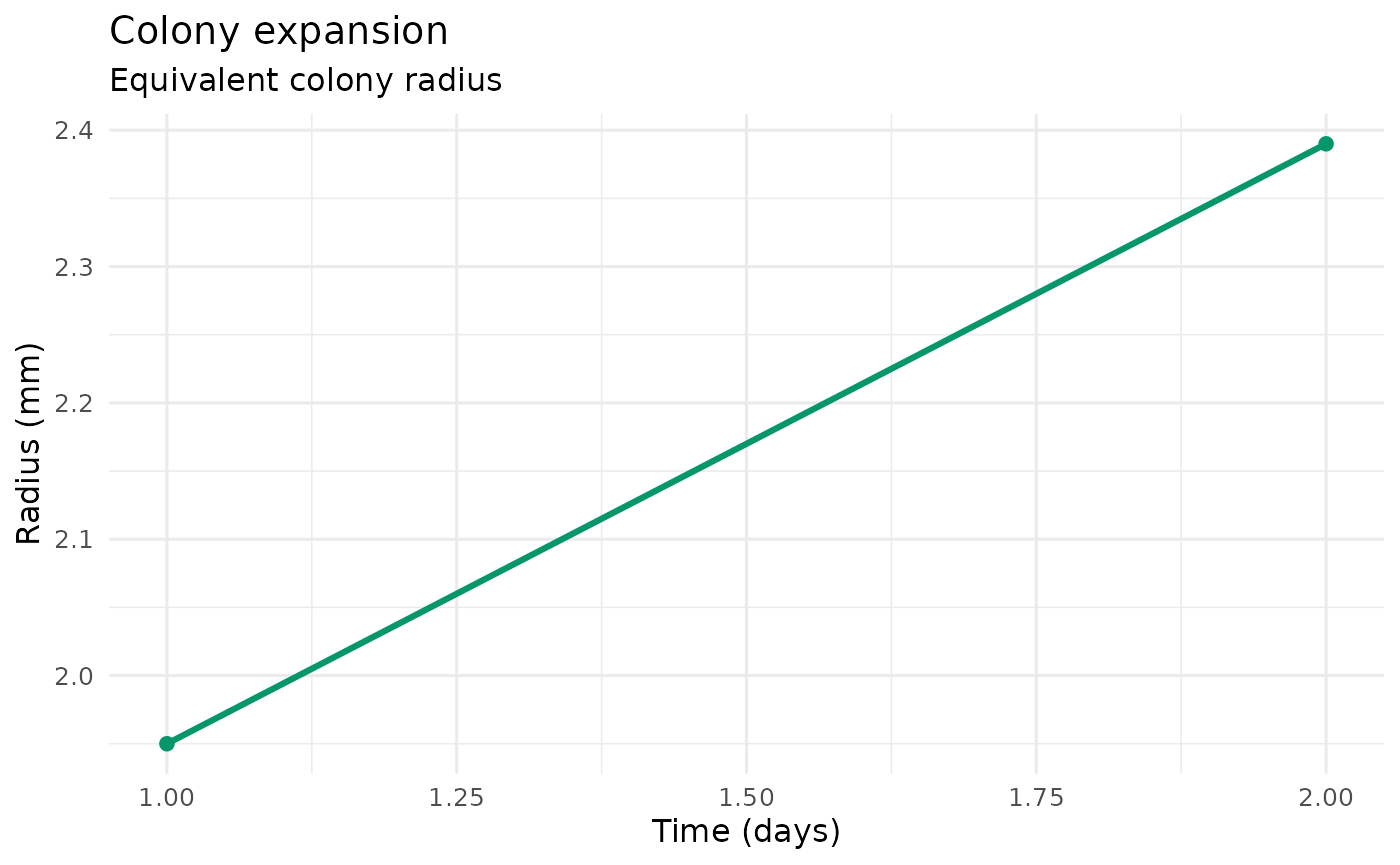
plot_growth_roughness(example_run)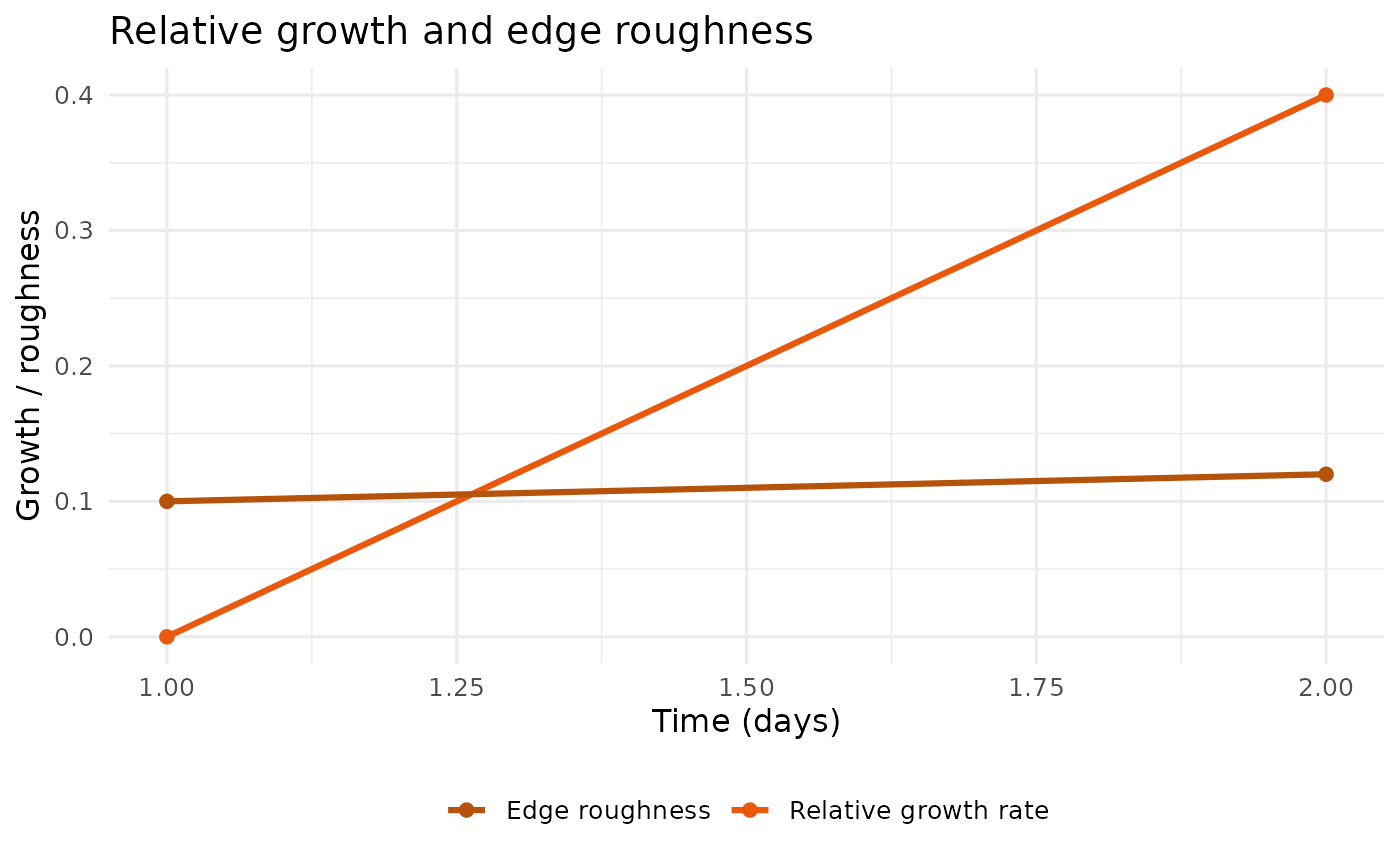
plot_stress_remodeling(example_run)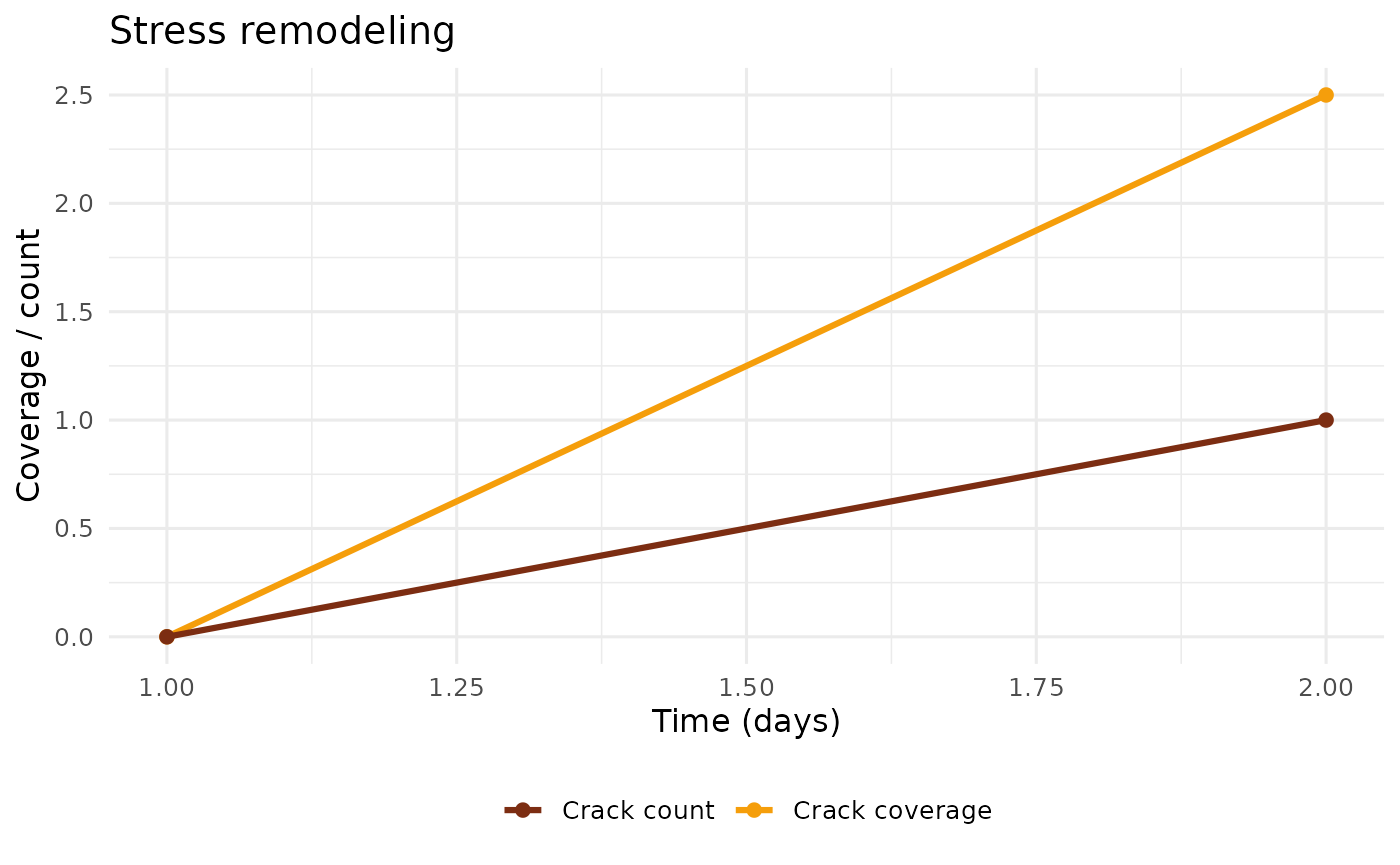
plot_texture_organization(example_run)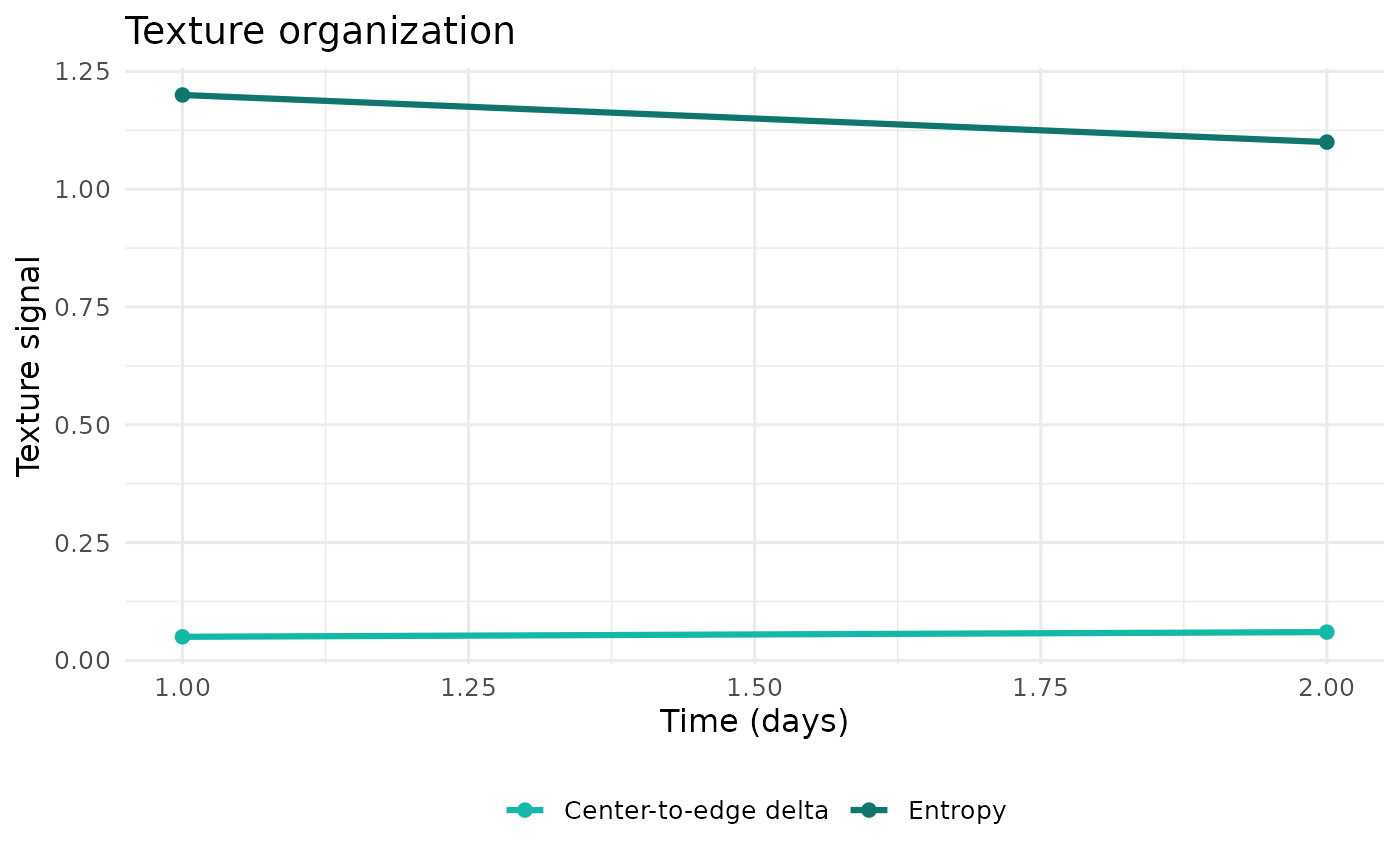
plot_shape_vs_stress(example_run)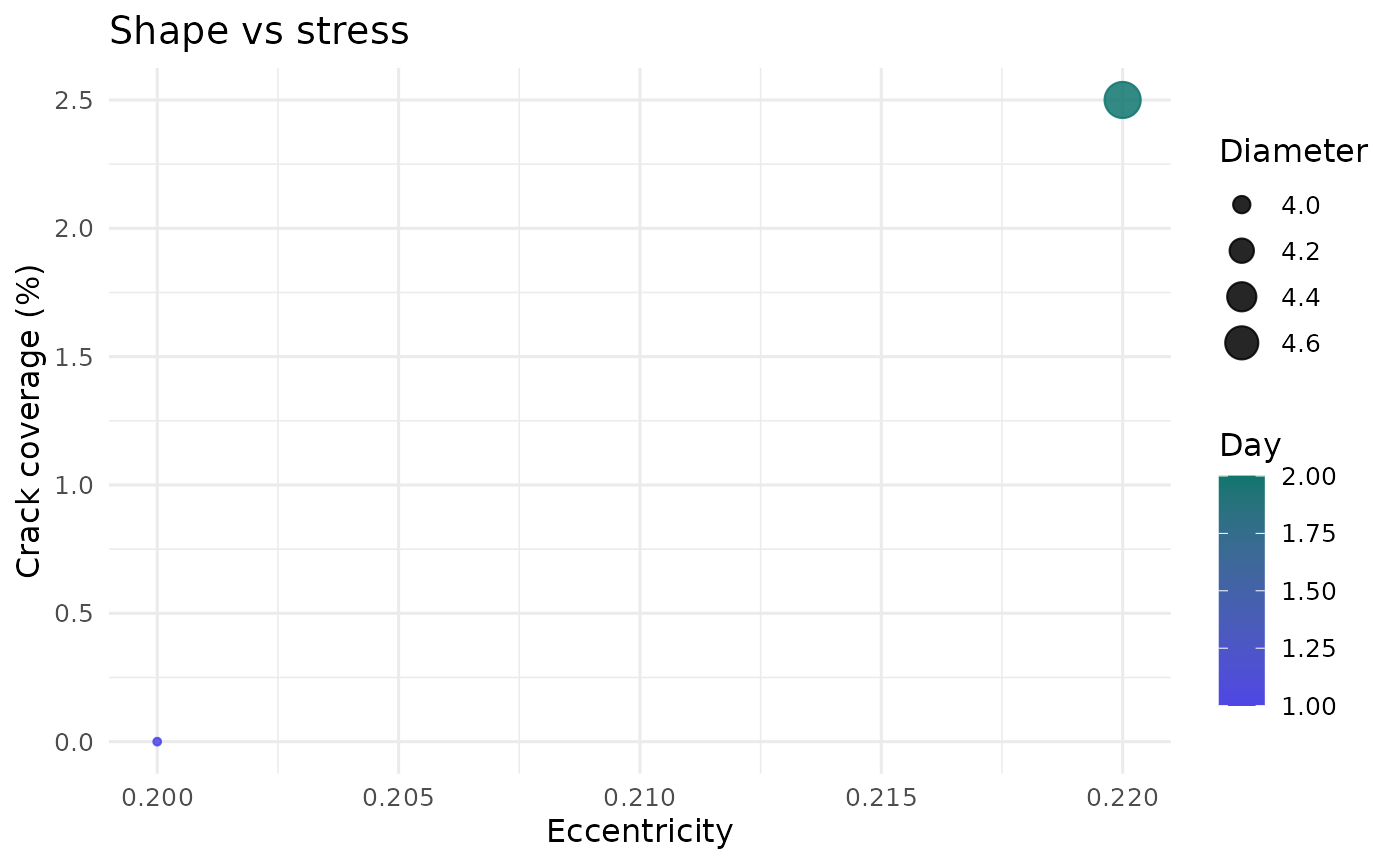
plot_feature_heatmap(example_run)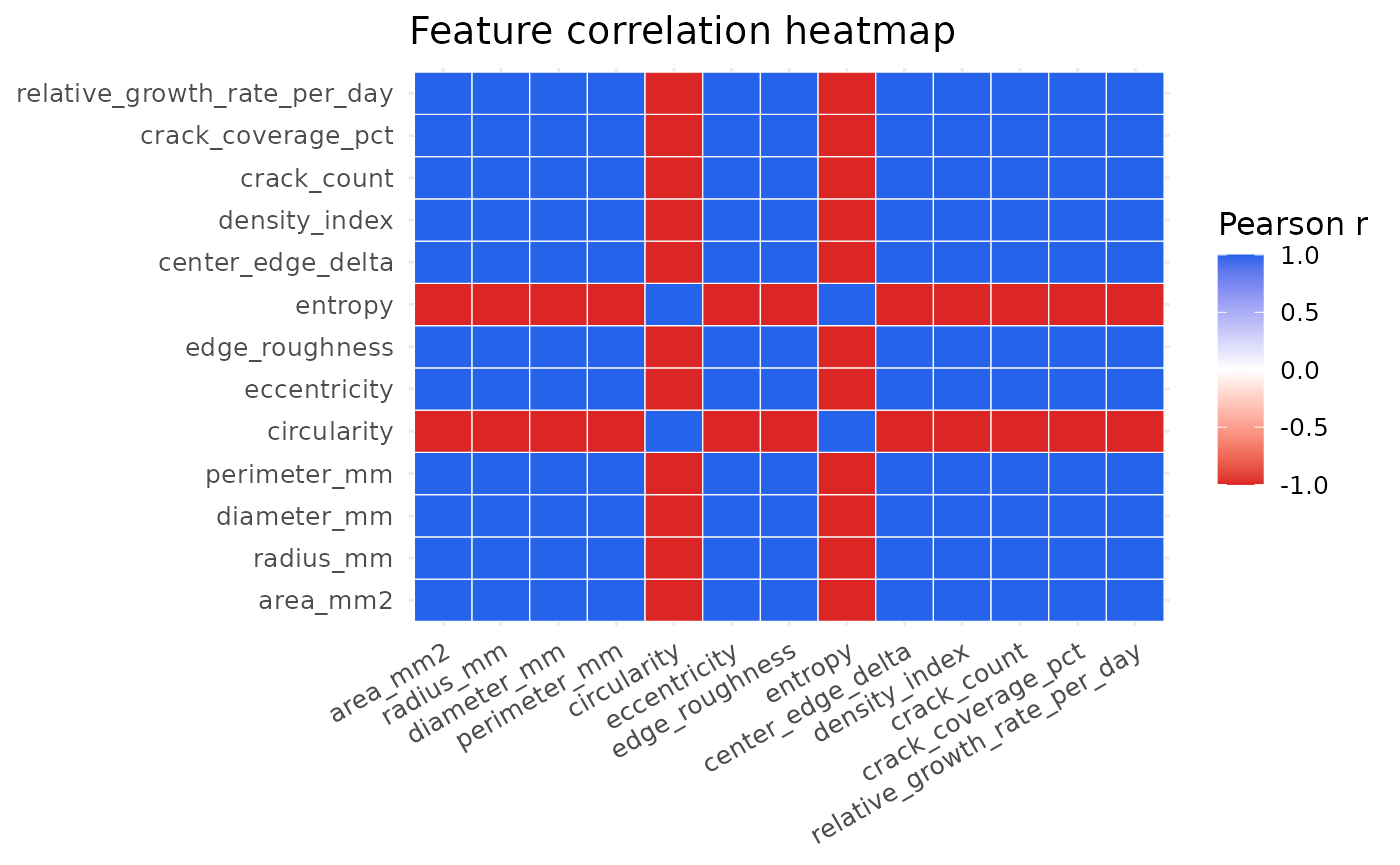
plot_radial_profile(example_run)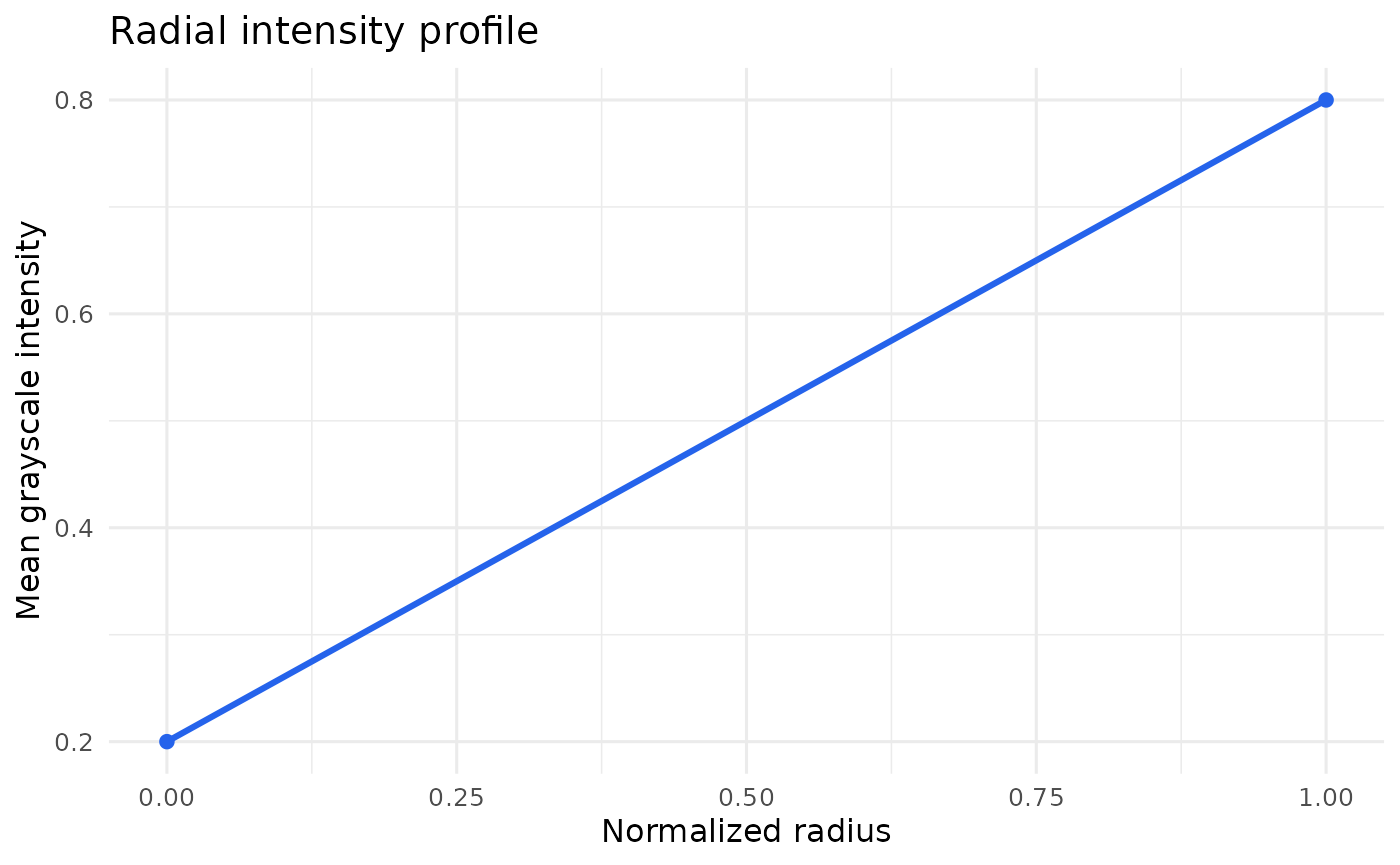
5. Run the Python analysis pipeline on real images
One-time Python setup
Add the interpreter path to ~/.Rprofile so it loads
automatically every time R starts:
file.edit("~/.Rprofile")
# Add this line (adjust path to your rvenv_arm_311 location):
# Sys.setenv(GRAYLEAFSPOTR_PYTHON = "/path/to/rvenv_arm_311/bin/python")Or set it for the current session only:
Sys.setenv(GRAYLEAFSPOTR_PYTHON = "/path/to/rvenv_arm_311/bin/python")Run with grayleafspot_run() — the primary entry
point
grayleafspot_run() finds the installed package Python
directory automatically, creates the output folder if needed, and
returns parsed JSON results directly.
res <- grayleafspot_run(
input_dir = system.file("extdata", "testdata", "06FEB", package = "grayleafspotr"),
output_dir = file.path(tempdir(), "gls_run"),
run_name = "trial_01"
)
# Access results
res$run # manifest metadata
res$results # per-image data frameFull-featured alternative: grayleafspot_analyze()
For tidy S3 objects with built-in plotting helpers, use
grayleafspot_analyze():
real_run <- grayleafspot_analyze(
input_dir = system.file("extdata", "testdata", "06FEB", package = "grayleafspotr"),
output_dir = tempdir(),
save_outputs = FALSE,
verbose = FALSE,
engine_model = "localunet"
)
plot_colony_expansion(real_run)6. Convert to tidy data
growth_data <- as_grayleafspot_growth_data(example_run)
growth_data## # A tibble: 2 × 34
## id filename day area_mm2 radius_mm diameter_mm perimeter_mm circularity
## <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 demo-1 demo_day… 1 12 1.95 3.9 8 0.8
## 2 demo-2 demo_day… 2 18 2.39 4.78 9.1 0.77
## # ℹ 26 more variables: eccentricity <dbl>, edge_roughness <dbl>,
## # contrast <dbl>, correlation <dbl>, energy <dbl>, homogeneity <dbl>,
## # entropy <dbl>, center_edge_delta <dbl>, density_index <dbl>, core <dbl>,
## # middle <dbl>, outer <dbl>, crack_count <dbl>, crack_length_mm <dbl>,
## # crack_coverage_pct <dbl>, proportional_crack_coverage_pct <dbl>,
## # radial_velocity_mm_per_day <dbl>, area_growth_rate_mm2_per_day <dbl>,
## # relative_growth_rate_per_day <dbl>, radial_acceleration <dbl>, …7. Custom plotting
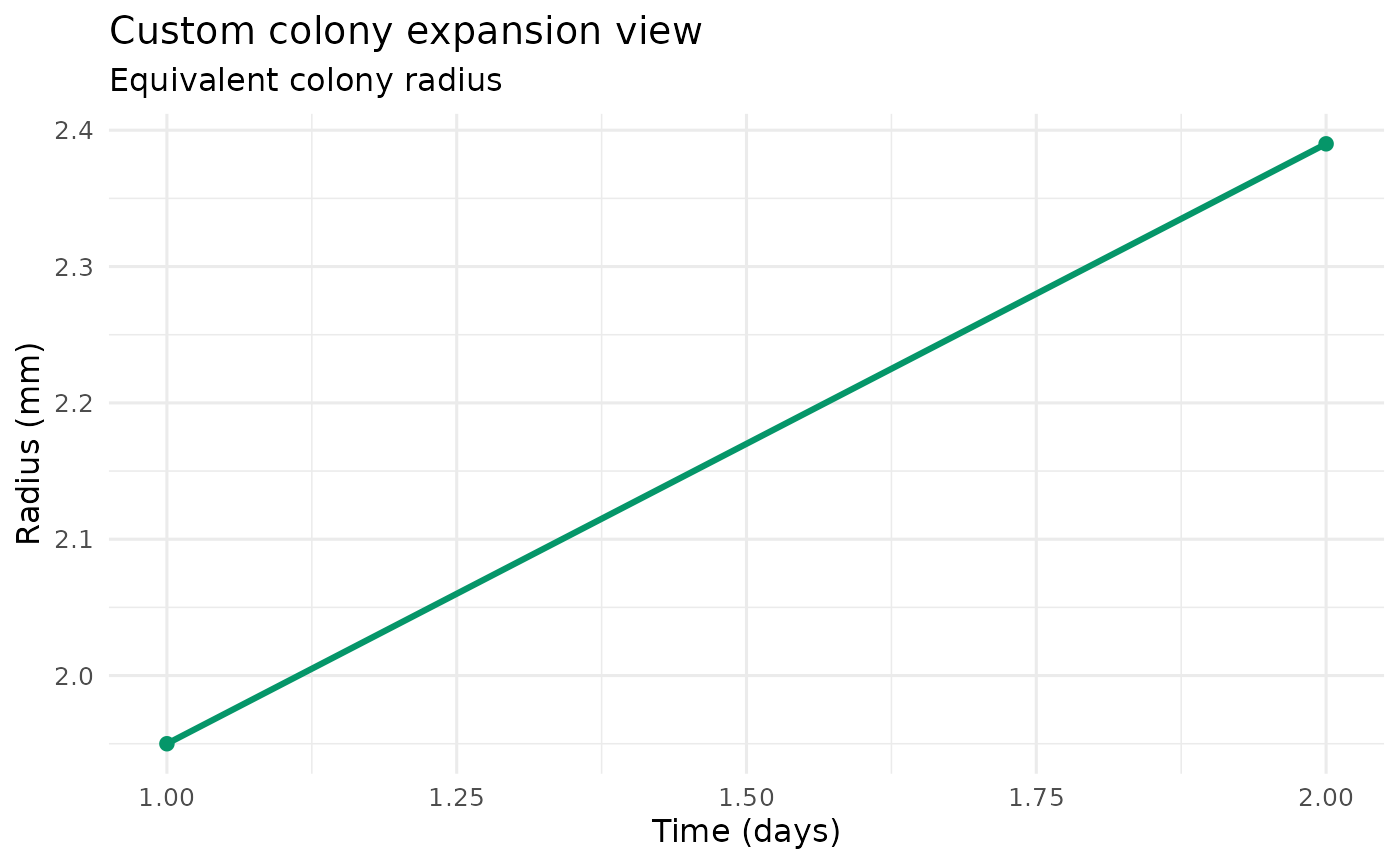
plot_colony_expansion(example_run) +
ggplot2::labs(title = "Custom colony expansion view")