Protein → RNA → DNA Simulator
A web application for exploring the inverted central dogma concept behind DRT3 protein-templated DNA synthesis.
This project is simulation and hypothetical only and has not been tested or validated. This project is for educational purposes only.
Features
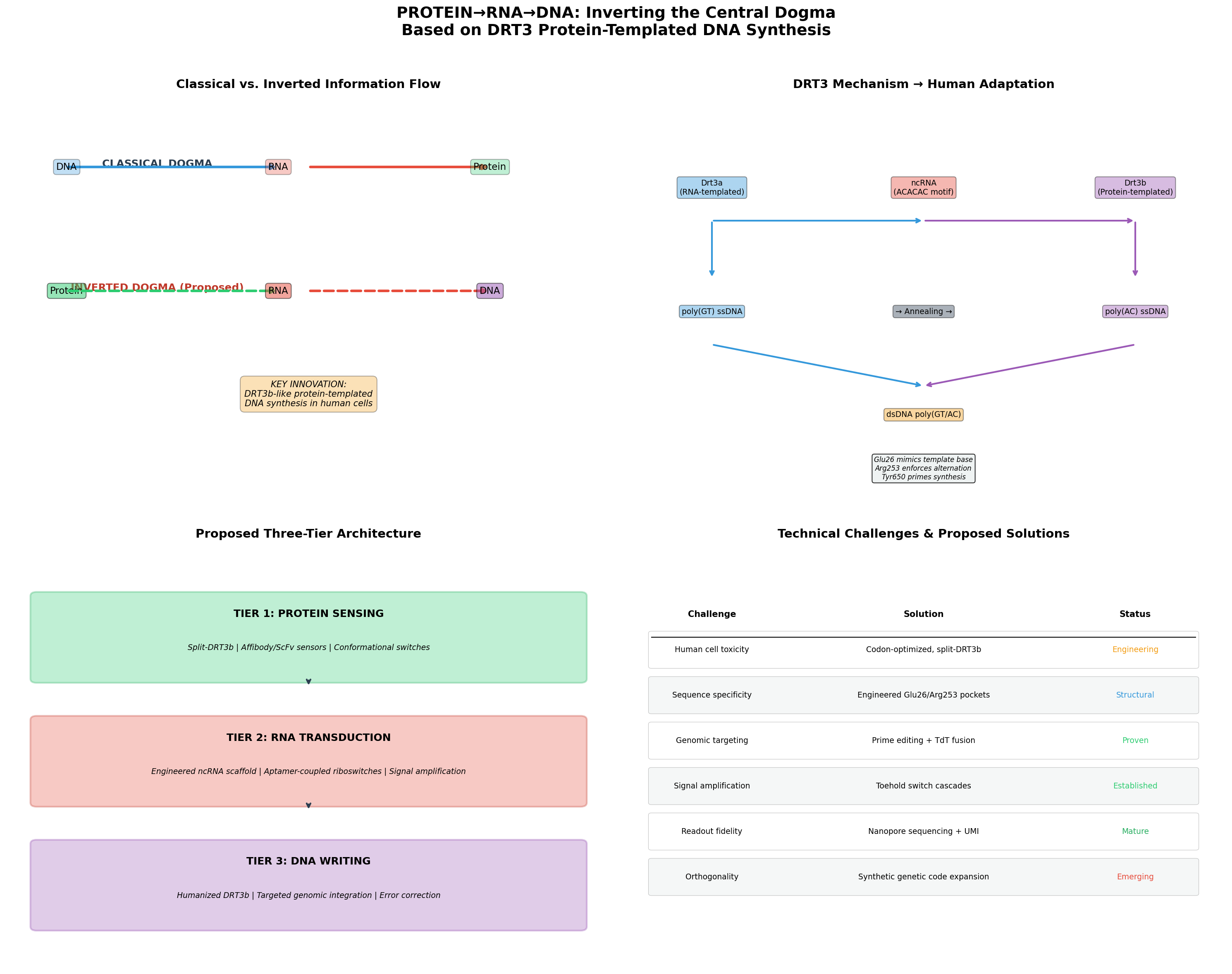
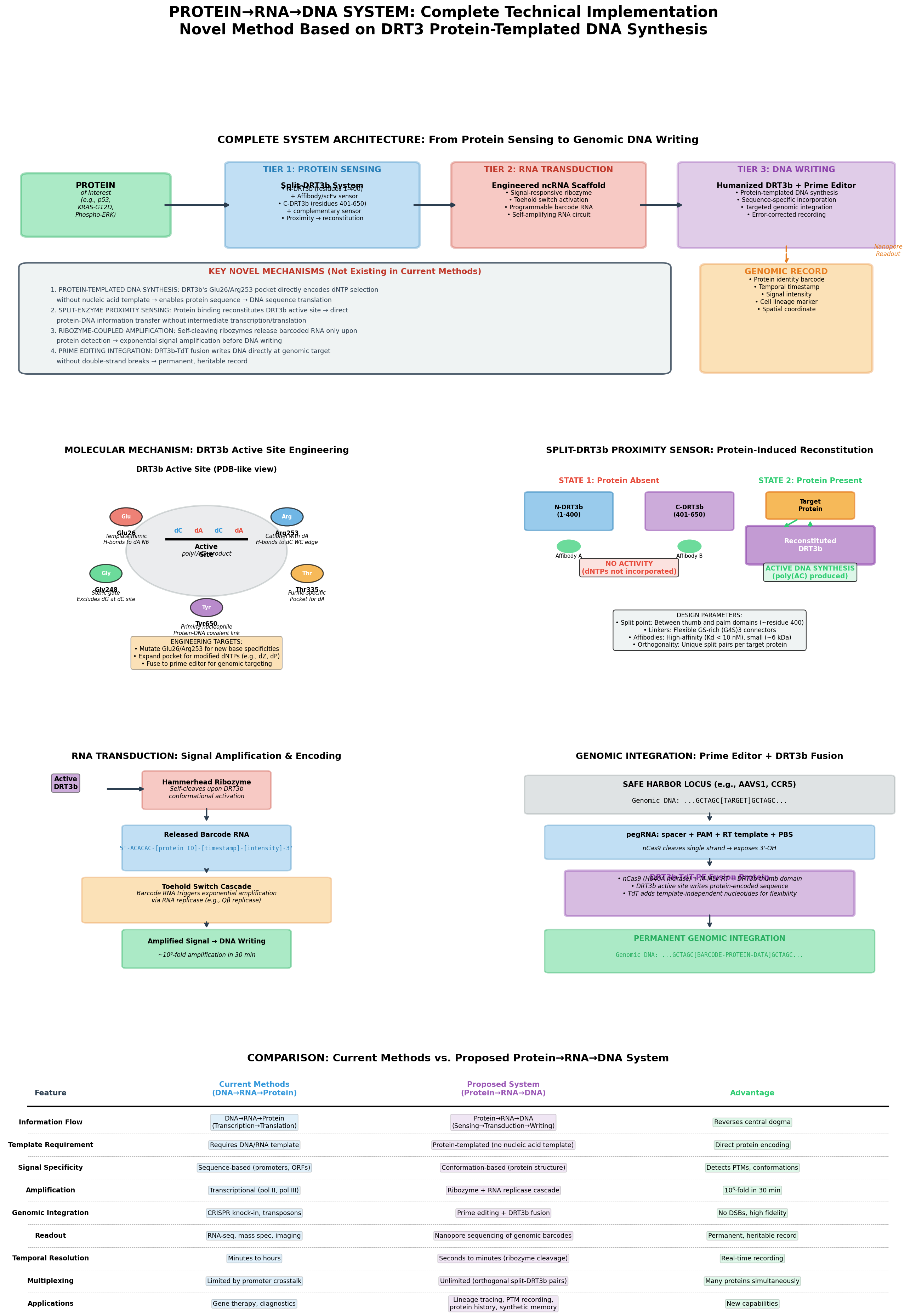
- Three-tier simulation dashboard (protein sensing → RNA amplification → DNA writing)
- Interactive 3D molecular viewer (NGL, PDB + AlphaFold integration)
- Curated protein browser with live AlphaFold DB and RCSB PDB fetching
- DRT3b active-site engineering lab (variant comparison, base-preference charts)
- Classical vs inverted central dogma comparison table
Scientific background and the full technical write-up live in protein_analysis.md.
Quick Start
1. Install dependencies
bun install
macOS Intel / Rosetta note: the system Node.js may run as x64 while Bun is arm64. If CSS stops rendering after
bun install, run once:
bash npm install lightningcss-darwin-x64 --no-save
2. Start the app
bun run dev
Then open http://localhost:3000.
Production build
bun run build
bun run start
Data Sources
| Source | Used for |
|---|---|
| RCSB PDB | Experimentally determined structures |
| AlphaFold DB | Predicted structures via UniProt accession |
| UniProt | Protein and gene cross-references |
Fetched protein data and dashboard state are stored in browser localStorage — the UI survives page refreshes without a database.
License
MIT License — copyright © 2026 Rohan R
Figures